Details of DPV and References
DPV NO: 211 July 1979
Family: Betaflexiviridae
Genus: Carlavirus
Species: Pea streak virus | Acronym: PeSV
This virus is now regarded as a distinct strain of pea streak virus
Alfalfa latent virus
V. Veerisetty Department of Plant Pathology, University of Missouri, Columbia, Missouri 65211, USA
Contents
- Introduction
- Main Diseases
- Geographical Distribution
- Host Range and Symptomatology
- Strains
- Transmission by Vectors
- Transmission through Seed
- Transmission by Grafting
- Transmission by Dodder
- Serology
- Nucleic Acid Hybridization
- Relationships
- Stability in Sap
- Purification
- Properties of Particles
- Particle Structure
- Particle Composition
- Properties of Infective Nucleic Acid
- Molecular Structure
- Genome Properties
- Satellite
- Relations with Cells and Tissues
- Ecology and Control
- Notes
- Acknowledgements
- Figures
- References
Introduction
- Described by Veerisetty & Brakke (1977a).
- An RNA-containing virus with straight to slightly flexuous particles about 635 nm long. Manually transmissible to a few species, mostly in the Leguminosae. Aphids transmit the virus in the non-persistent manner readily to pea and broad bean but with difficulty to alfalfa. Found in the USA.
Main Diseases
Occurs symptomlessly in alfalfa (Medicago sativa cv. Dawson).
Geographical Distribution
Recorded in the USA. Prevalent in Nebraska and Wisconsin.
Host Range and Symptomatology
Host range is very limited. Experimentally transmitted mainly to legumes. Alfalfa and hairy vetch (Vicia villosa) become infected without producing symptoms at 20 or 26°C. Pea (Pisum sativum cv. Lincoln) shows obvious symptoms only at 13°C.
- Diagnostic species
- Vicia faba
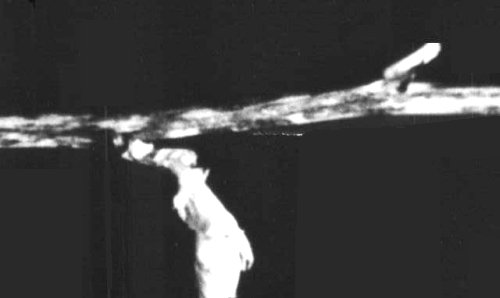
(broad bean). Very few dark reddish-brown necrotic local lesions appear on inoculated leaves followed by dark reddish-brown necrosis on top leaves. Wilting and abscission of the leaves with occasional dark reddish-brown necrosis on stems leading to the death of plants (Fig. 1). - Pisum sativum (pea) cv. Lincoln. Symptomless systemic infection at 20
or 26°C but at 13°C the plants show vein clearing, chlorosis of young leaves
and, more than 2 months after inoculation, necrosis of old leaves (Veerisetty &
Brakke, 1977a).
- Cassia occidentalis. The Wisconsin isolate induces small local lesions in inoculated leaves (Khan et al., 1977).
- Cassia occidentalis. The Wisconsin isolate induces small local lesions in inoculated leaves (Khan et al., 1977).
- Propagation species
- Pisum sativum.
Dwarf pea varieties (Lincoln and Little Marvel) are better than tall varieties (Tall Telephone and Alaska). Systemically infected Lincoln pea plants are a good source of virus for purification (Veerisetty & Brakke, 1978).- Assay species
- V. faba
is not a good assay host, though it reacts with local lesions; C. occidentalis may prove better (numerous local lesions are produced after inoculation with the Wisconsin isolate).
Strains
No critical tests were made to establish differences between Nebraska and Wisconsin isolates.
Transmission by Vectors
Pea aphids (Acyrthosiphon pisum), starved for 30 min and then allowed an acquisition access period of 5 to 10 min, transmitted the virus efficiently to pea and broad bean, regardless of virus source. When pea served as the virus source plant, the virus was consistently transmitted to alfalfa, but with low efficiency (Veerisetty & Brakke, 1977a).
Transmission through Seed
Not seed-transmitted in pea (Veerisetty & Brakke, 1977a).
Serology
The virus is moderately immunogenic. Antiserum with a precipitin titre of 1/512 was prepared by injecting rabbits (twice intramuscularly and once intravenously) with partially purified virus (10 mg total virus). The antiserum reacted with homologous antigen in microprecipitin tests, or in immunodiffusion tests in the presence of 0.05% sodium dodecyl sulphate.
Relationships
An isolate from Wisconsin is serologically related to the original isolate from Nebraska (Khan et al., 1977). Properties in vitro and particle morphology suggest that the virus should be placed in the carlavirus group but no serological relationship was detected to pea streak virus or red clover vein mosaic virus (Veerisetty & Brakke, 1977a).
Stability in Sap
In crude sap of Vicia faba, infectivity was lost after 4-6 days at 25°C, 10 min at 65-70°C, and the dilution end-point was between 10-3 and 10-4 (Veerisetty & Brakke, 1977a).
Purification
The following procedure consistently gives yields of about 0.7 to 1.0 mg virus/g tissue (Veerisetty & Brakke, 1978). Harvest pea cv. Lincoln tissue (whole plant excluding roots) 3 weeks after inoculation. Blend frozen tissue with two parts (w/v) of 0.165 M disodium phosphate + 0.018 M trisodium citrate buffer, pH 9.0, containing 0.1% diethyldithiocarbamate (DIECA) and 0.5% mercaptoethanol (ME) and centrifuge at low speed. Clarify the supernatant fluid by adding 0.05 vol. 0.2 M Na2HPO4 and 0.01 vol. 1.0 M CaCl2, stirring for 15-20 min and then centrifuging at low speed. Precipitate the virus from the supernatant fraction by dissolving polyethylene glycol (PEG, M. Wt 6000) to 6% (w/v) and centrifuging at low speed. Resuspend in 16 mM disodium phosphate + 2 mM trisodium citrate buffer, pH 9.0, containing 1% Triton X-100. Purify further by centrifuging the virus once each through 20 and 30% sucrose ‘cushions’.
Properties of Particles
Purified preparations contain a single infective nucleoprotein component when centrifuged in rate zonal sucrose or equilibrium CsCl density gradients. Sedimentation coefficient (s20,w): 161 ± 1.5 S (Veerisetty & Brakke, 1978).
A260/A280: 1.15-1.25; Amax(260)/Amin(244): 1.12 (both without correction for light-scattering).
M. Wt (obtained by calculation from M. Wt of RNA and protein subunits and
percentage RNA composition) is c. 4.7 x 107
(Veerisetty & Brakke, 1977b).
Particle Structure
Particles are straight to slightly flexuous filaments (Fig. 2), of helical construction. Modal lengths of c. 635 nm and 653 nm, respectively, were obtained with diffraction grating replica (22,835 lines/cm) and tobacco mosaic virus (300 nm) as standards (Veerisetty & Brakke, 1977a). Particles contain c. 1635 identical protein subunits packed in a helix (c. 8.2 subunits/turn). There are 5 nucleotides per subunit and the RNA is located at a radius of c. 3.3 nm from the centre of the particle (Veerisetty, 1978; Veerisetty & Brakke, 1977b).
Particle Composition
Nucleic acid: RNA, single-stranded, infective. Single component, M. Wt = 2.45 x 106 (estimated by sucrose density gradient sedimentation and by gel electrophoresis); s20,w = 38 S, probably 5.4% of particle weight as in other carlaviruses (Veerisetty & Brakke, 1977b).
Protein: A single type of protein is found by electrophoresis in polyacrylamide/SDS gels, with M. Wt c. 2.7 x 104 (Veerisetty & Brakke, 1977b).
Relations with Cells and Tissues
No inclusion bodies were found by light microscopy in epidermal strips of leaves, petioles and stems of infected pea and broad bean (Veerisetty & Brakke, 1977a).
Notes
Although particle morphology places alfalfa latent virus in the carlavirus group, it differs in host range and symptomatology from three other carlaviruses occurring naturally in legumes, namely cowpea mild mottle virus, pea streak virus, and red clover vein mosaic virus. Cowpea mild mottle virus does not infect Lincoln pea whereas alfalfa latent virus gives symptomless systemic infection; pea streak virus produces chlorosis and necrosis of the leaves and extensive stem necrosis, resulting in death of the plants; and red clover vein mosaic virus produces vein-clearing and chlorosis, curling, and downward rolling of the young leaves leading to terminal rosetting, stunting, and death of the plants. Alfalfa latent virus does not infect Trifolium pratense, T. repens or T. hybridum, which are hosts of red clover vein mosaic and pea streak viruses (Hagedorn & Hanson, 1951; Hagedorn & Walker, 1949; Veerisetty & Brakke, 1977a). Cowpea mild mottle virus does not infect Vicia faba but is easily transmitted to Vigna unguiculata and Glycine max whereas the reverse is true of alfalfa latent virus. Also cowpea mild mottle virus differs in not being transmitted by the pea aphid (Brunt & Kenten, 1973; Veerisetty & Brakke, 1977a).
Alfalfa latent virus is serologically unrelated to red clover vein mosaic and
pea streak viruses, and it also differs from all other legume carlaviruses in
coat protein M. Wt (Veerisetty & Brakke, 1977b). Alfalfa latent
virus should not be confused with lucerne latent virus (Taylor & Smith,
1971; Blackstock, 1978) which is a member of the nepovirus group, and has
been renamed lucerne Australian latent virus (Jones, Forster & Mohamed, 1979).
Figures
References list for DPV: Alfalfa latent virus (211)
- Blackstock, Aust. J. agric. Res. 29: 291, 1978.
- Brunt & Kenten, Ann, appl. Biol. 74: 67, 1973.
- Hagedorn & Hanson, Phytopathology 41: 813, 1951.
- Hagedorn & Walker, Phytopathology 39: 837, 1949.
- Khan, Maxwell, Palmer & Hanson, Proc. Am. Phytopath. Soc. 4: 122, 1977.
- Jones, Forster & Mohamed, Ann. appl. Biol. 92: 49, 1979.
- Taylor & Smith, Rep. Vict. Pl. Res. Inst. 5: 17, 1971.
- Veerisetty, Virology 84: 523, 1978.
- Veerisetty & Brakke, Phytopathology 67: 1202, 1977a.
- Veerisetty & Brakke, Virology 83: 226, 1977b.
- Veerisetty & Brakke, Phytopathology 68: 59, 1978.