Details of DPV and References
DPV NO: 297 September 1985
Family: Geminiviridae
Genus: Begomovirus
Species: African cassava mosaic virus | Acronym: ACMV
African cassava mosaic virus
K. R. Bock ICRISAT Regional Groundnut Program, Chitedze Research Station, Private Bag 63, Lilongwe, Malawi
B. D. Harrison Scottish Crop Research Institute, Invergowrie, Dundee DD2 5DA, UK
Contents
- Introduction
- Main Diseases
- Geographical Distribution
- Host Range and Symptomatology
- Strains
- Transmission by Vectors
- Transmission through Seed
- Transmission by Grafting
- Transmission by Dodder
- Serology
- Nucleic Acid Hybridization
- Relationships
- Stability in Sap
- Purification
- Properties of Particles
- Particle Structure
- Particle Composition
- Properties of Infective Nucleic Acid
- Molecular Structure
- Genome Properties
- Satellite
- Relations with Cells and Tissues
- Ecology and Control
- Notes
- Acknowledgements
- Figures
- References
Introduction
- Disease described by Warburg (1894) and Storey & Nichols (1938); virus
described by Bock et al. (1978) and Harrison et al. (1977).
- Synonyms
- Cassava latent virus (Rev. Pl. Path. 58: 1603)
- Cassava mosaic virus (Rev. appl. Mycol. 18: 230)
- Cassava mosaic virus (Rev. appl. Mycol. 18: 230)
- A virus with a genome of two circular molecules of single-stranded DNA contained in separate geminate particles of c. 30 x 20 nm. Transmitted in the persistent manner by the whitefly Bemisia tabaci and by inoculation with sap. Most reported natural hosts are in the Euphorbiaceae; experimental hosts include several solanaceous species. Occurs in Africa, nearby islands and India, and is of great economic importance.
Main Diseases
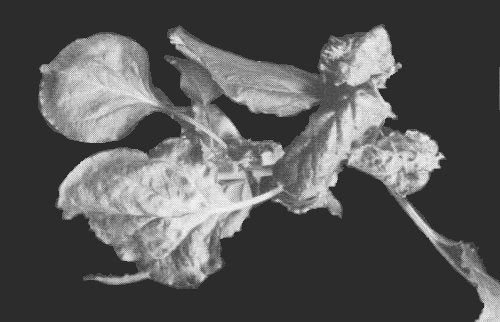
Causes a severe mosaic disease of cassava (Manihot esculenta, Euphorbiaceae; Fig. 1; Storey & Nichols, 1938; Bock & Woods, 1983). There is a 60-80% decrease in tuber yield of plants of susceptible varieties when infected cuttings are used for propagation (Bock, 1983). In many parts of Africa, the incidence of infection in cassava crops exceeds 80%, and the virus is of great economic importance.
Geographical Distribution
Occurs throughout Africa and Malagasy, in Indian Ocean Islands (Seychelles, Zanzibar, Pemba) and in India. The virus almost certainly originated in Africa (it has not been found in the New World), from where it has been disseminated to other areas, presumably in infected cassava cuttings.
Host Range and Symptomatology
Manihot esculenta is the only known naturally infected host of the Kenya type strain. The Kenya coast strain occurs naturally not only in M. esculenta but also in Jatropha multifida (Euphorbiaceae) and, apparently, in Hewittia sublobata (Convolvulaceae; Bock et al., 1981). In Nigeria, Laportea (= Fluerya) aestuans (Urticaceae) is suspected to be a natural host (Anon., 1979). The experimental host range of the Kenya type strain (as determined by inoculation with sap) is narrow and largely restricted to the Solanaceae, within which the virus is more or less readily transmitted to several species in the genera Nicotiana (benthamiana, clevelandii, debneyi, glutinosa, rustica, tabacum) and Datura (ferox, stramonium). Dubern (1979) transmitted an Ivory Coast isolate by whiteflies from M. esculenta to six additional Manihot spp. but not to 181 other species in 30 plant families.
- Diagnostic species
- Manihot esculenta
(cassava). Severe systemic mosaic in susceptible cvs 3-5 weeks after inoculation by whiteflies or by grafting. Flushes of symptom-bearing leaves may alternate with leaves that show few or no symptoms. Symptoms may be mild, intermediate or severe, depending on the virus isolate (Storey & Nichols, 1938). Infected by inoculation of sap, but only with difficulty (Bock & Woods, 1983). - Nicotiana benthamiana. Pronounced diffuse chlorotic local lesions
followed by severe systemic leaf curling, crinkling and yellow blotching, with
rapid and progressive decrease in leaf size and internode length (Fig. 2).
- N. clevelandii. The type strain may induce local chlorotic lesions (but not invariably) in 8-14 days; it always causes severe systemic leaf curl and stunting (Fig. 3), with subsequent development in some leaves of coarse irregular yellow veinbanding and yellow areas. Isolates of the Kenya coast strain either infect only with difficulty, to produce somewhat less severe symptoms, or do not infect.
- Datura stramonium. The type strain induces chlorotic and necrotic local lesions (Fig. 4), systemic veinbanding and severe leaf curling and distortion.
- N. clevelandii. The type strain may induce local chlorotic lesions (but not invariably) in 8-14 days; it always causes severe systemic leaf curl and stunting (Fig. 3), with subsequent development in some leaves of coarse irregular yellow veinbanding and yellow areas. Isolates of the Kenya coast strain either infect only with difficulty, to produce somewhat less severe symptoms, or do not infect.
- Propagation species
- Isolates are best maintained in vegetatively propagated cassava. They may
also be maintained in mechanically inoculated N. benthamiana but the
severity of symptoms necessitates frequent sub-culturing. N. benthamiana
is the best source of virus particles for purification.
- Assay species
- Although local lesions are induced in several species and D. stramonium has been used as a local lesion host for the type strain (Robinson et al., 1984), none is entirely reliable for local lesion assay. Assays of the Kenya coast strain must be made by recording systemic infection in batches of N. benthamiana. Cassava seedlings or cuttings are suitable test plants for transmission tests with Bemisia tabaci.
Strains
Type strain, isolate 844 (Bock et al., 1978). Obtained from W. Kenya, where other isolates differ in severity of symptoms in cassava. Antigenically similar isolates occur in Nigeria and Angola (Bock et al., 1981; Sequeira & Harrison, 1982) and seem widespread in areas of eastern Africa west of the Rift Valley and in West Africa. These isolates may differ somewhat in severity of symptoms in test plants such as N. benthamiana.
Kenya coast (C) strain (Bock et al., 1981). Transmitted with difficulty from cassava to N. benthamiana and with great difficulty to N. clevelandii, in which symptoms are less severe than those of the type strain. Distinguishable from, but related to, the type strain in serological (Bock et al., 1981) and nucleic acid hybridization tests, and with a higher optimum temperature for accumulation in plants (Robinson et al., 1984). Isolates of the C strain differ in the severity of symptoms induced in cassava (Bock et al., 1981). In Kenya, the different geographical distribution of the type and C strains may reflect two paths of entry of cassava, one across Africa from the west (type strain) and the other (C strain) from offshore islands, such as Zanzibar and Malagasy, in the east (Purseglove, 1968).
Indian strain. Serologically distinguishable from both the type and C strains in gel diffusion tests (K. R. Bock, P. Shanta & V. G. Malathi, unpublished results).
Angola defective isolates (Sequeira & Harrison, 1982). Not transmissible to N. benthamiana by inoculation with sap and apparently defective for particle production (Robinson et al., 1984). In cassava these isolates are graft transmissible and nearly all leaves on infected plants develop symptoms.
Transmission by Vectors
The virus is transmitted in the persistent manner by the whitefly Bemisia tabaci. Minimum acquisition access period is 3.5 h, minimum latent period is 8 h and minimum inoculation access period is 10 min (Chant, 1958; Dubern, 1979). Whiteflies remain infective for c. 9 days. About 10 per cent transmission can be obtained with single adult whiteflies. The virus also can be transmitted by the first three larval instars and is retained through the moult but is not passed through the egg to progeny insects (Dubern, 1979). In Kenya, spread of virus by whiteflies seems to occur mainly over short distances and long distance spread is uncommon (K. R. Bock & I. A. D. Robertson, unpublished results).
Transmission through Seed
The virus is not seed-borne in cassava (Storey & Nichols, 1938).
Transmission by Dodder
Not transmitted from cassava to cassava or other species by Cuscuta gronovii or C. subinclusa (Dubern, 1979).
Serology
Moderately immunogenic; rabbit antisera with titres of 1/500 in gel-diffusion precipitin tests with virus particles can be obtained using conventional injection schedules. These antisera have been used in precipitin tests (Bock et al., 1978), in ELISA and immunoelectron microscopy tests, in density-gradient zone depletion tests, and for fluorescent antibody staining (Sequeira & Harrison, 1982). In ELISA, antibodies to the type strain also detect the Kenya coast strain in cassava leaf extracts (A. M. Lennon, M. Aiton & B. D. Harrison, unpublished results).
Nucleic Acid Hybridization
Cloned probes have been prepared to each of the two genome DNA species (Stanley, 1983). Probes for either DNA species can be used in spot hybridization tests to detect infection of N. benthamiana or cassava with virus isolates resembling the type strain or the Angolan defective isolates, but only DNA-1 probes react strongly in tests for the Kenya coast strain in these two plant species (Robinson et al., 1984; D. J. Robinson & B. D. Harrison, unpublished results).
Relationships
The virus has a genome of circular, single-stranded DNA and a particle morphology typical for geminiviruses. Its particles are strongly serologically related to those of the following whitefly-transmitted members of the group: bean golden mosaic (Sequeira & Harrison, 1982), squash leaf curl (Cohen et al., 1983), tomato golden mosaic (Stein et al., 1983) and euphorbia mosaic (Roberts et al., 1984) viruses. In gel-diffusion precipitin tests, SDI values of about 2 are obtained in comparisons with bean golden mosaic and tomato golden mosaic viruses. In addition, nucleic acid hybridization tests detected sequence homologies between the larger genome part (DNA-1) of African cassava mosaic virus and the DNA of bean golden mosaic, tomato golden mosaic, tobacco leaf curl, tomato leaf curl and tomato yellow leaf curl viruses (Roberts et al., 1984). Within DNA-1 of African cassava mosaic virus, the sequence similarities with bean golden mosaic and tomato golden mosaic viruses are close in coding regions and scarcely evident in non-coding regions (Hamilton et al., 1984; Harrison, 1985). No serological relationship or genome homology has been confirmed between African cassava mosaic virus and any leafhopper-transmitted geminivirus.
Stability in Sap
Using the type strain, and N. clevelandii as source and assay host, the thermal inactivation point in sap is about 55°C, the dilution end-point about 10-3, and infectivity is decreased after 2-3 days at room temperature and lost after 3-4 days. Infectivity is retained for at least 6 months in frozen dehydrated leaves (Bock et al., 1978), or frozen leaves (J. C. Sequeira & B. D. Harrison, unpublished results).
Purification
(Sequeira & Harrison, 1982). Grow infected N. benthamiana at 20-25°C. Harvest systemically infected leaves 2-3 weeks after inoculation of the plants and store at -70°C for at least 2 days. Triturate frozen leaf tissue (100 g) in 200 ml 0.1 M Tris-HCl buffer, pH 8.4, containing 1% thioglycerol, and 100 ml chloroform. Centrifuge at 10,000 g for 10 min, then to the aqueous phase add polyethylene glycol M. Wt 6000 to 4% (w/v) and NaCl to 0.2 M. Stir for 2 h at 4°C, collect the precipitate and resuspend in 80 ml 0.01 M Tris-HCl, pH 8.0, containing 0.005 M sodium ethylenediamine-tetraacetate (TE buffer). Further purify by low- and high-speed centrifugation, resuspending the virus-containing pellets in 2 ml half-strength TE buffer. Clarify by low-speed centrifugation and sediment for 1 h at 250,000 g in 10-40% sucrose density gradients. Collect the light-scattering zone, dialyse overnight against TE buffer, concentrate the virus by high-speed centrifugation, resuspend it in 1 ml half-strength TE buffer and clarify. Yield can exceed 1 mg virus particles/100 g leaf.
Properties of Particles
Particles are stable in TE buffer but disintegrate in 0.03% sodium dodecyl
sulphate at pH 8.0, suggesting they are stabilized by protein-nucleic acid bonds
(Sequeira, 1982).
Sedimentation coefficient: s20,w= 76 S (main
component; Bock et al., 1978)
and c. 50 S (minor component).
Particle weight: 4.24 x 106 daltons (calculated from weights of
DNA and protein molecules).
A260/A280: 1.4 (Sequeira, 1982).
Particle Structure
Particles are geminate (c. 30 x 20 nm), with an obvious ‘waist’ at the mid-point of the long axis. Each half has an apparently pentagonal profile, with the faces in contact being longer than the others (Fig. 5). Preparations of purified particles also contain quasi-isometric particles c. 20 nm in diameter.
Particle Composition
Nucleic acid: Single-stranded circular DNA, about 22% of the particle weight and of M. Wt 0.92 x 106, estimated by electron microscopy (Fig. 6; Harrison et al., 1977; J. C. Sequeira, G. H. Duncan & B. D. Harrison, unpublished results). Nucleotide sequencing (Stanley & Gay, 1983) shows that there are two genome components, containing respectively 2779 residues (DNA-1; M. Wt 0.93 x 106) and 2724 residues (DNA-2; M. Wt 0.91 x 106). Each geminate particle accommodates one molecule of either DNA-1 or DNA-2. Some quasi-isometric particles of the type strain contain a smaller circular DNA of M. Wt c. 0.4 x 106 (Sequeira et al., 1983), which shares nucleotide sequences with DNA-2 (Stanley, 1983). Preparations of virus DNA contain, in addition to these circular molecules, linear molecules which share nucleotide sequences with DNA-1 and DNA-2, and which have a predominant size of about 0.8 x 106 daltons.
Protein: Particles contain one protein species comprising 78% of the particle weight. Its M. Wt was estimated by polyacrylamide gel electrophoresis in earlier work to be 34,000 (Bock et al., 1977) and more recently to be 31,800 ± 700 (Sequeira, 1982). The nucleotide sequence of the particle protein gene gives a M. Wt of 30,200 (Stanley & Gay, 1983). The protein does not contain detectable carbohydrate (Sequeira, 1982).
Genome Properties
Isolated virus DNA is about 2% as infective as the DNA in virus particles
(J. C. Sequeira & B. D. Harrison, unpublished results). Tests with cloned
double-stranded copies of DNA of the type strain show that both DNA-1 and DNA-2
are needed for infection of N. benthamiana, in which geminate particles
and both DNA species were produced but not the circular DNA molecules of
0.4 x 106 M. Wt (Stanley, 1983). The two genomic DNA species have
different sequences except for a shared ‘common region’ of 193 nucleotides.
In both genome parts, open reading frames occur both in the strand found in the
virus particles (+ strand) and in the complementary strand (- strand),
implying that transcription is bidirectional. Six of the open reading frames have
counterparts in the genomes of bean golden mosaic and tomato golden mosaic viruses,
and there is some overlap between open reading frames in different phases (Fig. 9).
Sequences that may act as promoters for transcription, and others that may act as
transcript termination and polyadenylation signals, are available for all the
above six open reading frames (Fig. 9). The common sequence contains a stem and
loop structure of 33 nucleotides which is a possible origin for DNA synthesis.
Pseudo-recombinants can be produced by reassorting the genome parts of two
parental isolates: DNA-1 determines coat protein specificity and the symptoms
in tobacco (Stanley et al., 1985).
Infected N. benthamiana leaves contain three double-stranded forms of genome DNA: circular molecules covalently closed in both strands, circular molecules covalently closed in one strand, and linear molecules (Stanley & Townsend, 1985). In addition, a double-stranded form was found of the small circular DNA molecule that occurs in plants infected with the type strain (D. J. Robinson, G. H. Duncan, J. C. Sequeira & B. D. Harrison, unpublished results). Polyadenylated RNA from infected plants includes transcripts of the + and - strands of both genome parts. The transcripts are of five sizes (0.7 to 1.7 kb) and can be tentatively assigned to the six shaded open reading frames in Fig. 9: a 1.0 kb transcript is the mRNA for the virus particle protein (Townsend et al., 1985).
Relations with Cells and Tissues
In N. benthamiana, virus particle antigen accumulates in the nuclei of many phloem parenchyma or companion cells and of some cells in other tissues, such as cortex, mesophyll and epidermis (Fig. 7 and 8; Sequeira & Harrison, 1982). Abnormalities in the nuclei of phloem parenchyma cells include the production both of granular inclusions that contain virus particles and of fibrillar rings (Horvat & Verhoyen, 1981; Adejare & Coutts, 1982). Similar abnormalities occur in cassava but are less common (Horvat & Verhoyen, 1981). Virus-free cassava plants can be produced from infected ones by heat treatment and meristem tip culture (Chant, 1959; Kaiser & Louie, 1982). Systemic invasion may be incomplete in plants of some cassava genotypes with resistance to the virus (H. Rossel, personal communication).
Notes
African cassava mosaic virus is the only geminivirus reported from
cassava. However, similar symptoms are caused by cassava common mosaic virus
(potexvirus group), which has been found only in South America, and also
differs in causing local lesions in Chenopodium amaranticolor and
Gomphrena globosa and in not being
transmitted by Bemisia tabaci (Costa & Kitajima, 1972). Viruses
associated with two other mosaic diseases of cassava in Colombia are not
sap-transmissible to N. benthamiana (A. M. Lennon, M. Aiton &
B. D. Harrison, unpublished results).
The close serological relationships between biologically distinct
geminiviruses can lead to difficulties in virus identification, especially
with infections of plant species not previously recorded as hosts. However,
unlike bean golden mosaic and squash mosaic viruses respectively, African
cassava mosaic virus does not infect Phaseolus vulgaris or
Cucurbita pepo. Also, in spot hybridization tests, probes for African
cassava mosaic virus DNA-2 do not react detectably with extracts of plants
infected with bean golden mosaic, tomato golden mosaic or tobacco leaf curl
and allied geminiviruses (Roberts et al., 1984).
Two methods of control of African cassava mosaic virus are advocated. In Kenya (Bock & Guthrie, 1982; Bock, 1983) and the Ivory Coast (Fargette et al., 1985), control can be achieved by propagating virus-free planting material of existing cultivars that have moderate resistance to infection. In Nigeria, control by breeding cultivars with superior resistance to, and tolerance of, infection is favoured (Hahn et al., 1980).
Acknowledgements
Figs 1-6 courtesy of the Scottish Crop Research Institute; Figs 7 and 8 from Sequeira & Harrison, 1982; Fig. 9 by courtesy of R. Townsend.
Figures

Symptoms in leaf of cassava (Manihot esculenta) cv. 46106/27 naturally infected with the Kenya coast strain.

Purified virus particles stained with uranyl acetate. Bar represents 50 µm. (Courtesy I. M. Roberts.)
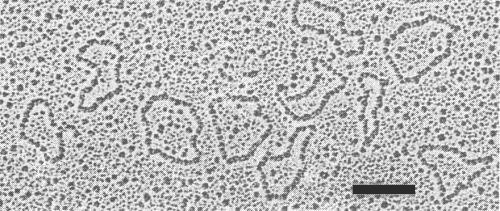
DNA molecules extracted from purified virus particles and rotary shadowed with uranium. Bar represents 200 nm. (Courtesy G. H. Duncan.)

Section of infected N. benthamiana leaf tissue stained with fluorescent antibody to the virus particles and examined (Fig. 7) in transmitted light and (Fig. 8) by fluorescence microscopy. Note nuclear staining in some cortical and epidermal cells. Bar represents 50 µm.
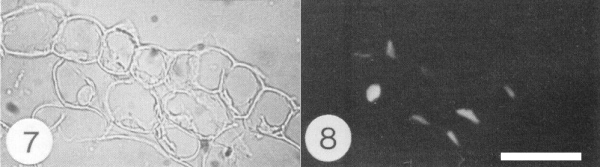
Section of infected N. benthamiana leaf tissue stained with fluorescent antibody to the virus particles and examined (Fig.7) in transmitted light and (Fig.8) by fluorescence microscopy. Note nuclear staining in some cortical and epidermal cells. Bar represents 50 µm.

Genome organization of African cassava mosaic virus (Townsend et al., 1985). Open reading frames are shown both in the strand in the virus particles (+ strand) and in the complementary strand (- strand) of DNA-1 (1) and DNA-2 (2). Open reading frames with counterparts in bean golden mosaic and tomato golden mosaic viruses are shaded. Figures indicate the M. Wt (x l0-3) of putative gene products. Nucleotides 1-193 comprise the common sequence. Possible transcription initiation and polyadenylation signals are indicated by open and closed triangles, respectively.
References list for DPV: African cassava mosaic virus (297)
- Adejare & Coutts, Phytopath. Z. 103: 87, 1982.
- Anon., Rep. int. lnst. trop. Agric. 1978: 107, 1979.
- Bock, in Plant Virus Epidemiology, p. 337, eds R. T. Plumb & J. M. Thresh. Oxford: Blackwell Scientific Publications, 377 pp., 1983.
- Bock & Guthrie, Trop. Pest Management 28: 219, 1982.
- Bock & Woods, Pl. Dis. 67: 994, 1983.
- Bock, Guthrie, Meredith & Barker, Ann. appl. Biol. 85: 305, 1977.
- Bock, Guthrie & Meredith. Ann. appl. Biol. 90: 361, 1978.
- Bock, Guthrie & Figueiredo. Ann. appl. Biol. 99: 151, 1981.
- Chant, Ann. appl. Biol. 46: 210, 1958.
- Chant, Emp. J. exp. Agric. 27: 55, 1959.
- Cohen, Duffus, Larsen, Liu & Flock, Phytopathology 73: 1669, 1983.
- Costa & Kitajima, CMI/AAB Descr. Pl. Viruses, 90, 4 pp., 1972.
- Dubern, Phytopath. Z. 96: 25, 1979.
- Fargette, Fauquet & Thouvenel, Ann. appl. Biol. 106: 285, 1985.
- Hahn, Terry & Leuschner, Euphytica 29: 673, 1980.
- Hamilton, Stein, Coutts & Buck, EMBO Jl. 3: 2197, 1984.
- Harrison, A. Rev. Phytopath. 23: 55, 1985.
- Harrison, Barker, Bock, Guthrie, Meredith & Atkinson, Nature, Lond. 270: 760, 1977.
- Horvat & Verhoyen, Parasitica 37: 119, 1981.
- Kaiser & Louie, Pl. Dis. 66: 475, 1982.
- Purseglove, Tropical Crops, Dicotyledons, London: Longman, 1968.
- Roberts, Robinson & Harrison, J. gen. Virol. 65: 1723, 1984.
- Robinson, Harrison, Sequeira & Duncan, Ann. appl. Biol. 105: 483, 1984.
- Sequeira, Ph.D. Thesis, University of Dundee, 1982.
- Sequeira & Harrison, Ann. appl. Biol. 101: 33, 1982.
- Sequeira, Harrison & Duncan, Rept. Scott. Crop Res. Inst., 1982: 193, 1983.
- Stanley, Nature, Lond. 305: 643, 1983.
- Stanley & Gay, Nature, Lond. 301: 260, 1983.
- Stanley & Townsend, Nucleic Acids Res. 13: 2189, 1985.
- Stanley, Townsend & Curson, J. gen. Virol. 66: 1055, 1985.
- Stein, Coutts & Buck, J. gen. Virol. 64: 2493, 1983.
- Storey & Nichols, Ann. appl. Biol. 25: 790, 1938.
- Townsend, Stanley, Curson & Short, EMBO Jl. 4: 33, 1985.
- Warburg, Mitt. dt. Schutzgeb. 7: 131, 1894.