Details of DPV and References
DPV NO: 317 December 1986
Family: Unallocated ssRNA+ viruses
Genus: Sobemovirus
Species: Velvet tobacco mottle virus | Acronym: VTMoV
Velvet tobacco mottle virus
J. W. Randles Waite Agricultural Research Institute, University of Adelaide, South Australia, Australia
R. I. B. Francki Waite Agricultural Research Institute, University of Adelaide, South Australia, Australia
Contents
- Introduction
- Main Diseases
- Geographical Distribution
- Host Range and Symptomatology
- Strains
- Transmission by Vectors
- Transmission through Seed
- Transmission by Grafting
- Transmission by Dodder
- Serology
- Nucleic Acid Hybridization
- Relationships
- Stability in Sap
- Purification
- Properties of Particles
- Particle Structure
- Particle Composition
- Properties of Infective Nucleic Acid
- Molecular Structure
- Genome Properties
- Satellite
- Relations with Cells and Tissues
- Ecology and Control
- Notes
- Acknowledgements
- Figures
- References
Introduction
Virus purified and described by Randles et al. (1981).
A virus with c. 30 nm isometric particles containing a linear single-stranded RNA of M. Wt c. 1.5 x 106. All naturally occurring isolates studied also contain a circular single-stranded RNA of M. Wt 0.12 x 106, which is probably a satellite. The virus occurs in wild plants of Nicotiana velutina in south central Australia and infects a narrow range of experimental hosts. It is readily transmissible by mechanical inoculation of sap and its natural vector is the mirid bug, Cyrtopeltis nicotianae.
Main Diseases
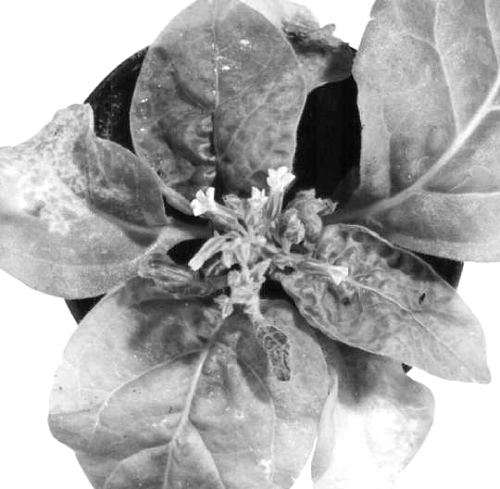
In Nicotiana velutina, the only known natural host, the virus (all naturally occurring isolates studied contain the satellite RNA) causes systemic vein clearing followed by mosaic with leaf distortion (Randles et al., 1981; Fig. 1).
Geographical Distribution
The virus has been recovered only from N. velutina plants growing near Cobblers Sandhill and Innamincka in north-eastern South Australia (Randles et al., 1981) and near Cunnamulla in south-western Queensland (G. Behncken & J. W. Randles, unpublished data). The distribution of N. velutina in Australia (Horton, 1981) may determine the distribution of the virus.
Host Range and Symptomatology
Transmitted readily by sap inoculation to a few species in the Solanaceae (Francki et al., 1985).
-
Diagnostic species
- Nicotiana clevelandii.
Inoculated leaves develop chlorotic to necrotic local lesions followed by systemic vein-yellowing and mosaic (Fig. 3). An experimentally obtained isolate lacking the small circular RNA (Francki et al., 1986) produced no obvious lesions and a milder mosaic. - N. glutinosa and N. edwardsoni (N. clevelandii x N. glutinosa hybrid).
Mild mosaic.
- N. velutina. Systemic vein-yellowing and mosaic (Fig. 2).
-
Propagation species
- N. clevelandii
is used for maintenance and propagation of virus, and for culturing the mirid vector.Assay species
- N. clevelandii
is a local lesion host (Fig. 3) for cultures that contain the small circular RNA. - N. clevelandii is also used as a systemic host in vector studies.
Strains
No strains detected.
Transmission by Vectors
Cyrtopeltis nicotianae (Hemiptera : Miridae; Fig. 6) is considered to be the vector of velvet tobacco mottle virus in the field (Randles et al., 1981), but experimentally the virus is also transmitted by the coccinellid vectors (Epilachna spp.) of solanum nodiflorum mottle virus (R. Greber & J. W. Randles, unpublished data). Both nymphs and adults of C. nicotianae transmit the virus (J. W. Randles, unpublished data); they have an acquisition threshold of approximately 1 min, and an inoculation threshold of 2 h after a 24-48 h acquisition period. No latent period has been detected. The bugs continue to transmit the virus for up to 9 days, but transmit it intermittently when transferred daily to new test plants. Trans-stadial transmission from nymph to nymph and from nymph to adult has been demonstrated but no multiplication of the virus has been detected in the vector (K. S. Gibb & J. W. Randles, unpublished data).
Transmission through Seed
No seed transmission has been detected in N. velutina (Randles et al., 1981).
Serology
Strongly immunogenic; rabbit antisera had homologous titres of 1/1024 in double diffusion tests in 0.75% agar buffered with 10 mM phosphate, pH 7 (Randles et al., 1981; Chu & Francki, 1983).
Relationships
Velvet tobacco mottle virus is serologically related to solanum nodiflorum mottle virus, with a serological differentiation index of 1-2 (Randles et al., 1981; Chu & Francki, 1983). However, the two viruses differ in host range, notably in that S. nodiflorum is infected by solanum nodiflorum mottle virus but not by velvet tobacco mottle virus, whereas N. velutina and N. glutinosa are infected systemically by velvet tobacco mottle virus, but not by solanum nodiflorum mottle virus (Randles et al., 1981; Greber, 1981). Moreover, there is only 20% homology between the genomic RNA molecules (RNA1) of the two viruses (Gould & Hatta, 1981); however, the RNA2 molecule of velvet tobacco mottle virus has extensive sequence homology (92-95%) with that of solanum nodiflorum mottle virus, although it is 11-12 nucleotides smaller (Gould & Hatta, 1981; Haseloff & Symons, 1982).
Velvet tobacco mottle virus is also serologically unrelated to two other viruses that have icosahedral particles containing long linear and small circular RNA molecules, lucerne transient streak and subterranean clover mottle (Tien et al ., 1981; Francki et al., 1983b). The RNA2 molecules of lucerne transient streak and solanum nodiflorum mottle viruses behave as satellite RNA (Jones & Mayo, 1983; Paliwal, 1984), and so it now seems does that of velvet tobacco mottle virus (Francki et al., 1986 - see below). Moreover, although the RNA2 molecules of velvet tobacco mottle and solanum nodiflorum mottle viruses differ in size and sequence from those of lucerne transient streak and subterranean clover mottle viruses (Keese et al., 1983; Francki et al., 1985), they are replicated in association with lucerne transient streak virus (Jones & Mayo, 1983, 1985). Furthermore, the RNA2 molecules of lucerne transient streak virus are replicated in association with southern bean mosaic virus (Paliwal, 1984). These observations suggest that there are affinities among all these viruses and between them and the sobemoviruses.
Velvet tobacco mottle virus also resembles the sobemoviruses in particle morphology (Tremaine & Hamilton, 1983). However, no serological cross-reactions have been detected in gel double diffusion tests between velvet tobacco mottle virus and southern bean mosaic virus or any of the following viruses with small polyhedral particles sedimenting as a single component: broad bean mottle, carnation mottle, carnation ringspot, cymbidium ringspot, galinsoga mosaic, glycine mottle, narcissus tip necrosis, red clover necrotic mosaic, saguaro cactus, tomato bushy stunt and turnip crinkle viruses (M. Hollings, personal communication; Randles et al., 1981; Francki et al., 1983a).
Stability in Sap
The virus is infective in N. clevelandii sap after heating for 10 min at 70°C.
Purification
A suitable method (Randles et al., 1981) is to blend infected leaf material with 2 vol (w/v) 70 mM phosphate buffer, pH 7, containing 3 mM EDTA and 0.1% thioglycollic acid. After straining through cheesecloth, clarify the extract with 9% n-butanol at 40° for 20 min, centrifuge at 10,000 g for 10 min, then concentrate the virus particles from the low speed supernatant fluid by centrifugation at 78,000 g for 4 h. Resuspend the pellets in 70 mM phosphate buffer, pH 7, clarify by low speed centrifugation, and centrifuge the supernatant fluid at 200,000 g for 75 min. Resuspend the pellets in buffer and emulsify with an equal volume of chloroform. Collect the aqueous phase after centrifugation and centrifuge again at high speed. Purify the virus further by sucrose density gradient centrifugation in 5-25% sucrose gradients buffered in 20 mM phosphate buffer, pH 7.4 (Beckman SW 41 rotor, 40,000 rev/min for 1 h).
Properties of Particles
Virus particle preparations contain a single sedimenting component co-sedimenting with southern bean mosaic virus, which has a s20,w of 115 S (Tremaine & Hamilton, 1983).
Absorption coefficient (A260(0.1%, 1 cm)): approximately 5.
A260/A280: 1.54.
Buoyant density in CsCl: 1.37 g/cm3 (Randles et al., 1981).
Particle Structure
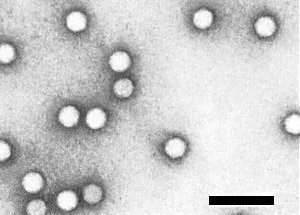
Negatively stained particles are icosahedral, about 30 nm in diameter (Fig. 7).
Particle Composition
Nucleic acid: Preparations of the virus particles contain linear single-stranded RNA of about 4500 nucleotides (RNA1), and two less abundant apparently subgenomic RNA components (RNA1a and RNA1b) of M. Wt 0.63 x 106 and 0.25 x 106 respectively (Gould, 1981). Particle preparations normally also contain two low M. Wt RNA species: covalently linked circular molecules (RNA2; Fig. 4) and molecules of identical size and sequence, but linear (RNA3; Fig. 4). Their M. Wt is 0.12 x 106, and they have no sequence homology with RNA1 (Randles et al., 1981; Gould, 1981). The RNA content of the particles is about 20%, and estimates of the relative amounts of the five RNA species are 7% (1) 40 (1a), 7% (1b), 47% (2) and 35% (3) (Randles et al., 1981). No particle heterogeneity has been observed, and it therefore seems likely that all particles contain the same weight of RNA, made up of either a single molecule of RNA1 or approximately 12 molecules of RNA2 and/or RNA3.
RNA2 contains 366 or 367 nucleotides; the sequence (Haseloff & Symons, 1982; Keese & Symons, 1987; Fig. 8) indicates regions of base-pairing that allow the molecule to adopt a secondary structure like that of viroids. RNA2 has no extensive sequence homology with viroids (Haseloff & Symons, 1982), but it exhibits co-operative thermal denaturation kinetics similar to that obtained for viroids but with a lower Tm of 38° in 10 mM NaCl, 1 mM Na cacodylate, 0.1 mM EDTA (Randles et al., 1982). Its circularity can be demonstrated by electron microscopy (Randles et al., 1981; Fig. 4) or by two-dimensional polyacrylamide gel electrophoresis (Schumacher et al., 1983). RNA2 has a 2' phosphomonoester group with the core structure C(2'p)3'p5'A at residue number 49 in the sequence; this is the probable ligation site for circularization of the molecule (Kiberstis et al ., 1985).
Protein: Dissociated virus particles yield polypeptides of M. Wt 31,500, 33,000 and 37,000 (Randles et al., 1981; Chu & Francki, 1983). However, the native virus particles probably contain a single protein species of M. Wt 37,000, which is partially degraded to the smaller components during virus purification (Chu & Francki, 1983).
Genome Properties
Infectivity of a natural isolate of the virus was reported by Gould et al. (1981) to be dependent on both RNA1 and RNA2 (or RNA3); the function of RNA2 was not replaced by the structurally similar RNA2 from solanum nodiflorum mottle virus. However, an isolate of the virus obtained experimentally (K1) contains no detectable RNA2 (Francki et al., 1986) and is distinguishable from natural isolates by the production of a milder mosaic and the absence of necrotic local lesions in N. clevelandii. This isolate supports the synthesis of exogenous RNA2 and encapsidates it, indicating that the RNA2 functions as a satellite RNA.
Double-stranded forms of both RNA1 and RNA2 are detectable during virus replication in N. clevelandii (Chu et al., 1983).
Relations with Cells and Tissues
Virus particles, c. 22 nm in diameter, are detected in tissue following in situ ribonuclease treatment (Hatta & Francki, 1981). They are distributed in nuclei, cytoplasm and cytoplasmic vacuoles, but are more numerous in the cytoplasm (Randles et al., 1981; Fig. 5). Some infection-specific vesicles containing electron-dense strands occur in the endoplasmic reticulum and perinuclear spaces but are more numerous in the cytoplasm (Fig. 5).
Notes
The virus cross-reacts serologically with solanum nodiflorum mottle virus, but is readily distinguishable from it by its failure to infect Solanum nodiflorum, and by the smaller size of its associated RNA2 (Haseloff & Symons, 1982).
Figures

RNA species 1, 1a, 1b, 2 and 3 separated on a denaturing polyacrylamide gel; and rotary-shadowed molecules of RNA species 1, 2 and 3 spread after denaturation. Bar represents 500 nm.

Thin section of an infected leaf cell of N. clevelandii, following treatment with ribonuclease. Note virus particles in nucleus, cytoplasm and vacuole, and electron-dense strands (arrowed). Bar represents 500 nm.

Primary nucleotide sequence (J02439)and suggested secondary structure of the 366-nucleotide form of RNA2 (after Keese & Symons, 1986). Nucleotides are numbered clockwise from the G residue at the left-hand end.
References list for DPV: Velvet tobacco mottle virus (317)
- Chu & Francki, Virology 129: 350, 1983.
- Chu, Francki & Randles, Virology 126: 480, 1983.
- Francki, Chu & Keese, in Curr. Comm. Mol. Biol.: Plant Infectious Agents-Viruses,Viroids, Virusoids, and Satellites, pp. 175-80, ed. H.D. Robertson et al., Cold Spr. Harb. Lab., 1983a.
- Francki, Grivell & Gibb, Virology 148: 381, 1986.
- Francki, Randles, Chu, Rohozinski & Hatta, in Subviral Pathogens of Plants and Animals:Viroids and Prions, Chapter 13, pp. 265-297, ed. K. Maramorosch & J. J. McKelvey, New York:Academic Press, 1985.
- Francki, Randles, Hatta, Davies & Chu, Pl. Path. 32: 47, 1983b.
- Greber, Aust. J. biol. Sci. 34: 369, 1981.
- Gould, Virology 108: 123, 1981.
- Gould & Hatta, Virology 109: 137, 1981.
- Gould, Francki & Randles, Virology 110: 420, 1981.
- Haseloff & Symons, Nucleic Acids Res. 10: 3681, 1982.
- Hatta & Francki, J. Ultrastruct. Res. 74: 1, 1981.
- Horton, J. Adelaide bot. Gard. 3: 1, 1981.
- Jones & Mayo, J. gen. Virol. 64: 1771, 1983.
- Jones & Mayo, Rep. Scott. Crop Res. Inst., 1984: 190, 1985.
- Keese & Symons, in Viroids and Viroid-Like Pathogens, ed. J. S. Semancik, Boca Raton: CRC Press, 1987.
- Keese, Bruening & Symons, FEBS Lett. 159: 185, 1983.
- Kiberstis, Haseloff & Zimmern, EMBO Jl 4: 817, 1985.
- Paliwal, Can, J. Pl. Path. 6: 93, 1984.
- Randles, Davies, Hatta, Gould & Francki, Virology 108: 111, 1981.
- Randles, Steger & Riesner, Nucleic Acids Res. 10: 5569, 1982.
- Tien, Davies, Hatta & Francki, FEBS Lett. 132: 353, 1981.
- Schumacher, Randles & Riesner, Anal. Biochem. 135: 288, 1983.
- Tremaine & Hamilton, CMI/AAB Descr. Pl. Viruses 274, 6 pp., 1983.