Details of DPV and References
DPV NO: 397 June 2003
Family: Alphaflexiviridae
Genus: Allexivirus
Species: Shallot virus X | Acronym: ShVX
Shallot virus X
S.K. Zavriev All-Russia Institute of Agricultural Biotechnology, Moscow, 127550, Russia
V.K. Vishnichenko All-Russia Institute of Agricultural Biotechnology, Moscow, 127550, Russia
Contents
- Introduction
- Main Diseases
- Geographical Distribution
- Host Range and Symptomatology
- Strains
- Transmission by Vectors
- Transmission through Seed
- Transmission by Grafting
- Transmission by Dodder
- Serology
- Nucleic Acid Hybridization
- Relationships
- Stability in Sap
- Purification
- Properties of Particles
- Particle Structure
- Particle Composition
- Properties of Infective Nucleic Acid
- Molecular Structure
- Genome Properties
- Satellite
- Relations with Cells and Tissues
- Ecology and Control
- Notes
- Acknowledgements
- Figures
- References
Introduction
Described by Vishnichenko et al. (1993).
A virus with flexuous filamentous particles about 800 nm long. Transmissible by mites (Aceria tulipae) and by sap inoculation. Widespread in shallot (Allium ascalonicum).
Main Diseases
Occurs in shallot, apparently symptomlessly, but detailed comparison with virus-free plants has not been done.
Geographical Distribution
Initially identified and described in Russia (Vishnichenko et al., 1993, Kanyuka et al., 1992), and also observed in the Netherlands and Germany; probably occurs world-wide.
Host Range and Symptomatology
Host range extremely narrow. Has been experimentally transmitted to onion, where systemic infection was symptomless, and to Chenopodium murale, which reacted with local lesions (van Dijk et al., 1991).
- Diagnostic species
- Chenopodium murale. Clearly defined small rings, often with a necrotic center, developing in inoculated leaves 2-3 weeks after inoculation. Not systemic (Van Dijk et al., 1991).
- Propagation species
- Shallot (Allium ascalonicum) is suitable for maintaining cultures, but care must be taken to select plants that are not infected with other viruses (Vishnichenko et al., 1996). Shallot is also a good source of virus for purification (Vishnichenko et al., 1993).
- Assay species
- Chenopodium murale is suitable for local lesion assay. Onion and shallot seedlings are useful for testing transmission by vectors.
Strains
None reported.
Transmission by Vectors
The only vector known is the mite Aceria tulipae. To avoid mechanical transmission, thin slices of mite-infested bulbs, sprouts or leaves were placed inside a paper cone attached around test plants which were usually incubated for several days in a nontransparent bag before transfer to the glasshouse (van Dijk et al., 1991).
Serology
The virus seems highly immunogenic. An antiserum with a titre of 1.8x10-6 in indirect ELISA tests was obtained from a rabbit injected with as little as 200 µg of a CsCl-purified virus preparation (Vishnichenko, Arshava, unpublished results). A recombinant ShVX coat protein has also been used as an immunogen (Arshava et al., 1995).
Relationships
ShVX is the type member of the Genus Allexivirus. Poorly characterized viruses serologically related to ShVX have been found in several Allium species (van Dijk et al., 1991, 1994), and in tulip and narcissus plants, but it is unclear whether these should be regarded as strains of ShVX or as distinct viruses. The serological relationships of ShVX to other well characterized allexiviruses or to viruses in other genera have not been studied.
Purification
Vishnichenko et al. (1993). Shallot plants (2-3 weeks after bulb planting) were homogenized with mortar and pestle in buffer A: (0.1 M K-phosphate pH 8.2, 5 mM EDTA, 0.2% Na2SO3) at a 1:2 (w/v) ratio. The homogenate was filtered through muslin and the extract was centrifuged at 12,000g for 25 min. Triton X-100 was added to the supernatant to a final conc. of 1%. The mixture was incubated at 4°C for 15 h and then centrifuged at 17000g for 15 min. The virus was pelleted by centrifugation (rotor SW41, 40,000 rpm, 3 h, 12°C) through a 2 ml layer of sucrose solution with a specific gravity of 1.2 g.cm-3 prepared in buffer B (0.01 M K-phosphate pH 7.4, 5 mM EDTA). Pellets were resuspended in sterile water, and insoluble material was removed by centrifugation in an Eppendorf microfuge. Virus can be further purified in CsCl gradients.
Properties of Particles
S20,w is about 170S in 0.1 M Tris-HC1, pH 7.5 at 20°C.
Buoyant density in CsCl is 1.33 g.cm-3.
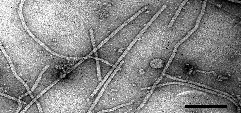
Particle Structure
Particles are flexuous filaments, about 800 nm long and 12 nm in diameter, helically constructed with a pitch of 2.5 nm (Fig. 1).
Particle Composition
Nucleic acids: Virions contain a single molecule of linear (+)-sense ssRNA, 8832 nt long, with a 3'-poly(A) tract. In RNA preparations from ShVX particles, besides genomic ssRNA, molecules of dsRNA, 1.5 kbp in length, are found whose structure, genesis and function(s) are unknown (Vishnichenko et al., 1993).
Proteins: Virions contain two serologically identical major coat proteins: one has a predicted Mr of 28K and the other an apparent Mr of 37K. Minor amounts of ORF4 encoded protein (Mr 42K) were also detected in the viral particles (Vishnichenko et al., 2002). The mechanism of the synthesis of the major coat proteins with different Mr from the same RNA is still obscure.
Genome Properties
The complete nucleotide sequence of the ShVX genomic RNA has been determined by Kanyuka et al. (1992); (GenBank accession number M97264).
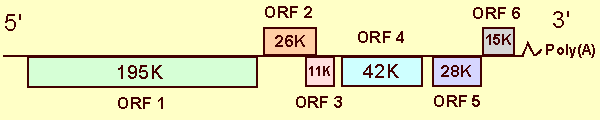
The genome organization of ShVX is shown in Fig. 2. The ShVX genomic RNA contains six large ORFs and non-coding sequences of 98 nts at the 5' terminus, and 112 nts followed by a poly(A) tail at the 3' terminus (Fig. 2). The ORFs code for polypeptides of Mr 195K, 26K, 11K, 42K, 28K and 15K, respectively, from the 5'- to the 3'-end. The gene arrangement of other sequenced allexiviruses is similar. The 195K polypeptide is probably the viral RNA polymerase. In comparisons among the amino acid sequences of methyltransferase, helicase, or polymerase domains, those of allexiviruses were most similar to those of potexviruses. The 26K and 11K proteins are similar to the first two proteins encoded by the triple gene block of potexviruses and carlaviruses and most probably are involved in cell-to-cell movement of the virus. There is a coding sequence for a small (Mr 7-8K) triple gene block protein but it lacks the AUG initiation codon. The 42K polypeptide has no significant homology with any proteins known, but has been shown to be expressed in plants infected with ShVX in relatively large amounts, and to be involved in virion assembly. The 28K polypeptide is the coat protein. In PAGE it migrates as a polypeptide with Mr of 32-36K, which could be due to the high hydrophilicity evident from its amino acid sequence. The 15K protein is similar to the 11-14K proteins encoded by the 3'-most ORFs of carlaviruses, has a zinc-binding finger motif and an ability to bind nucleic acids. The function of this polypeptide is not known.
Relations with Cells and Tissues
In partially purified shallot leaf extracts viewed in the electron microscope (immunodecoration test), the particles are usually present in high concentrations. They often occur in bundles, or in groups forming large amorphous aggregates. When viral infection is initiated in sprouting bulbs, synthesis of intact coat protein is observed in mitochondria, while the microsomal fraction contains short polypeptides serologically related to the coat protein. At later stages of infection, the capsid protein is synthesized mostly in the microsomal fraction. The unique allexiviral 42K protein is synthesized in microsomes at all stages of infection (Vishnichenko, unpublished results).
Notes
The virus (named Shallot mite-born latent virus) was probably first observed in the Netherlands, but was erroneously classified in the genus Rymovirus, family Potyviridae (van Dijk et al., 1991). This error has now been realized (van Dijk and van der Vlugt, 1994; Barg et al., 1994), and ShVX is considered as the type member of a new Genus Allexivirus (Zavriev et al., 2000).
Figures
References list for DPV: Shallot virus X (397)
- Arshava, Konareva, Ryabov & Zavriev, Molecular Biology (Russia) 29: 192, 1995.
- Barg, Lesemann, Vetten & Green, Acta Horticulturae 358: 251, 1994.
- Kanyuka, Vishnichenko, Levay, Kondrikov, Ryabov & Zavriev, Journal of General Virology 73: 2553, 1992.
- Van Dijk, Verbee & Bos, Netherlands Journal of Plant Pathology97: 381, 1991.
- Van Dijk & van der Vlugt, European Journal of Plant Pathology 100: 269, 1994.
- Vishnichenko, Konareva & Zavriev, Plant Pathology 42: 121, 1993.
- Vishnichenko, Stelmashchuk & Zavriev, Molecular Biology (Russia) 30: 959, 1996.
- Vishnichenko, Stelmashchuk & Zavriev, Molecular Biology (Russia) 36: 1080, 2002
- Zavriev, Ryabov & Vishnichenko, in Virus Taxonomy. 7th Report of the International Committee on Taxonomy of Viruses, p.981, eds M.H.V. van Regenmortel, C.M. Fauquet, D.H.L.Bishop et al., New York: Academic Press, 2000.