Details of DPV and References
DPV NO: 404 February 2004
Family: Bromoviridae
Genus: Ilarvirus
Species: Elm mottle virus | Acronym: EMoV
This is a revised version of DPV 139
Elm mottle virus
A.T. Jones Scottish Crop Research Institute, Invergowrie, Dundee DD2 5DA, UK
S.W. Scott Dept of Entomology, Soils, and Plant Sciences, Clemson University, Clemson, SC 29634-0315, USA
Contents
- Introduction
- Main Diseases
- Geographical Distribution
- Host Range and Symptomatology
- Strains
- Transmission by Vectors
- Transmission through Seed
- Transmission by Grafting
- Transmission by Dodder
- Serology
- Nucleic Acid Hybridization
- Relationships
- Stability in Sap
- Purification
- Properties of Particles
- Particle Structure
- Particle Composition
- Properties of Infective Nucleic Acid
- Molecular Structure
- Genome Properties
- Satellite
- Relations with Cells and Tissues
- Ecology and Control
- Notes
- Acknowledgements
- Figures
- References
Introduction
Described by Schmelzer et al. (1966) and Schmelzer (1969).
- Synonyms
- Lilac white mosaic virus (Schmelzer, 1969)
- Hydrangea mosaic virus (Scott et al., 2003)
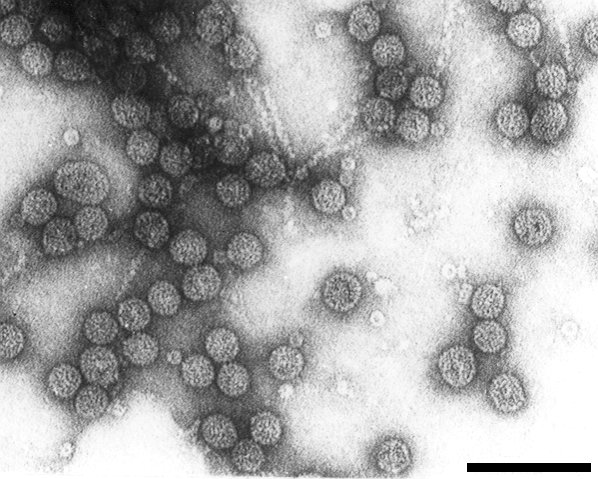
A virus with quasi-spherical particles c. 30 nm in diameter. Found in Europe. Virus particle preparations sediment as three nucleoprotein components which contain at least four ssRNA species of c. 3.43, 2.87, 2.31 and 1 kb, and a major protein of Mr24,000. Readily sap-transmissible to many herbaceous hosts. Seed-borne in elm. Vector not known.
Main Diseases
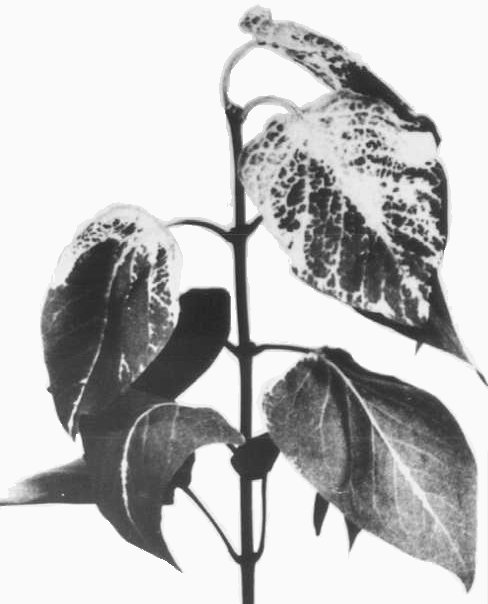
Induces a systemic white mosaic or chlorotic ringspots in mechanically inoculated Forsythia intermedia and Syringa vulgaris (Fig. 1) (Schmelzer, 1969). It is associated with ringspot and line-pattern leaf symptoms in elm (Fig. 2, Fig. 3) (Schmelzer, 1969; Jones & Mayo, 1973), but mechanically inoculated elm seedlings did not develop leaf symptoms when kept in a heated glasshouse (Jones & Mayo, 1973).
Geographical Distribution
Detected in Syringa vulgaris and Ulmus carpinifolia in Germany (Schmelzer, 1969) and in Ulmus glabra in Germany and Scotland (Schmelzer, 1969; Jones & Mayo, 1973). On the basis of leaf symptoms in elm it possibly also occurs in Bulgaria, Czechoslovakia, England and the former USSR (Schmelzer et al., 1966).
Host Range and Symptomatology
Found naturally infecting Ulmus carpinifolia, U. glabra and Syringa vulgaris (Schmelzer, 1969; Jones & Mayo, 1973). A German isolate infected 66 species from 22 families by mechanical inoculation (Schmelzer, 1969). These included many members of the Chenopodiaceae, Solanaceae and Leguminoseae. Many developed symptoms and most were infected systemically.
- Diagnostic species
- Chenopodium quinoa. Faint chlorotic local lesions; mottle or mosaic in systemically infected leaves in 7-10 days (Fig. 4).
- Phaseolus vulgaris (French bean) and Vigna sinensis (cowpea). Necrotic local lesions and small rings in 4-5 days (Fig. 5); not systemic.
- Nicotiana megalosiphon. Necrotic local lesions in 5 days (Fig. 6); systemic necrotic or chlorotic spots.
- Propagation species
- Chenopodium quinoa and Nicotiana tabacum (tobacco) have been used. Petunia hybrida is useful for maintaining cultures.
- Assay species
- Chenopodium quinoa appears the most sensitive indicator for detecting the virus from natural hosts. Phaseolus vulgaris and Vigna sinensis are excellent local lesion hosts.
Strains
Schmelzer (1969) distinguished isolates from Syringa vulgaris and Ulmus glabra by minor differences in host range and by antiserum titres. The Scottish isolate differs from the German isolates in being symptomless in Nicotiana clevelandii and Petunia hybrida (Jones & Mayo, 1973). The Scottish and German isolates are serologically related but it is not known whether they are serologically distinguishable (Jones & Mayo, 1973).
Transmission by Vectors
No vector reported. The aphids Aphis craccivora and Myzus persicae did not transmit the virus to herbaceous test plants after 15-30 min access periods on infected herbaceous plants (Schmelzer, 1969). Some viruses to which this virus is related are transmitted through infected pollen to pollinated plants but no critical tests to demonstrate this for EMoV have been done (Jones & Mayo, 1973).
Transmission through Seed
Seed-borne in Ulmus glabra to 20% (Jones & Mayo, 1973) and detected in the pollen of infected Syringa vulgaris (Schmelzer, 1969).
Transmission by Dodder
Not transmitted by either Cuscuta californica or C. subinclusa (Schmelzer, 1969).
Serology
The virus is weakly immunogenic, giving polyclonal antisera with precipitin end-points of 1/32-1/64. It reacts well with antiserum and produces a single line of precipitation in gel-diffusion tests.
Nucleic Acid Hybridization
cRNA probes to a fragment of the coat protein gene, detected by chemi-luminescent substrates, have been used successfully in dot blot and Northern hybridization procedures to detect virus in C. quinoa sap (S. W. Scott, unpublished data).
Relationships
The virus is serologically indistinguishable from Hydrangea mosaic virus (HdMV) (Jones, 1985) and the latter name should be considered a synonym of EMoV (Scott et al., 2003). EMoV is serologically related to Asparagus virus 2, Citrus leaf rugose virus, Citrus variegation virus and Tulare apple mosaic virus (Uyeda & Mink, 1983). These five viruses, together with Spinach latent virus, are in Subgroup 2 of the genus Ilarvirus in the family Bromoviridae (Roosinck et al., 2000). Viruses within this subgroup are related serologically but are not related to members of the other subgroups. Nucleotide sequence data confirm the relationships among members of subgroup 2 ( Scott et al., 1998, 2003).
Stability in Sap
All described isolates have similar properties (Schmelzer, 1969; Jones & Mayo, 1973). In C. quinoa sap, the Scottish isolate lost infectivity after 10 min at 55-60 °C, storage at room temperature for 9 days or dilution in 0.006 M phosphate buffer (pH 7.0) to 10-3-10-4. Sap kept at 4 °C or -15 °C was infective after 32 days, and leaves of C. quinoa freeze-dried by vacuum sublimation gave infective extracts after 2 years, the longest period tested (Jones & Mayo, 1973).
Purification
Using the Scottish isolate, Jones & Mayo (1973) obtained good yields of virus relatively free from host plant material by the following procedure. Extract inoculated and systemically infected leaves of C. quinoa in 0.05 M citrate buffer (pH 7.0). Squeeze the extract through muslin and clarify it by adding n-butanol (to 8.5% v/v) and stirring for 30-60 min. Concentrate and further purify the virus by differential centrifugation. Resuspend the virus-containing pellets from high speed centrifugation in 0.005 M citrate buffer (pH 7.0).
Properties of Particles
Purified preparations contain at least three nucleoprotein components distinguishable by their sedimentation rates (Fig. 7). Most of the infectivity is associated with the fastest sedimenting component (component 3), but it is enhanced by adding either, or both, of the two slower sedimenting components (components 1 and 2) (Jones & Mayo, 1973). No nucleic acid-free particles have been observed.
Sedimentation coefficients (s°20, w): 83 S (1), 88 S (2), 101 S (3) (Jones & Mayo, 1973).
A260/A280 of unfractionated virus: 1.3 (Jones & Mayo, 1973).
Buoyant density: when centrifuged to equilibrium in CsCl solution, unfractionated virus formed two bands of density c. 1.27 and c. 1.35 g.cm-3. Only the band of greater buoyant density was infective but both bands contained virus-like particles. However, whereas most of the particles in the infective band were c. 28 nm in diameter, most of those in the non-infective band were 15-22 nm in diameter (Jones & Mayo, 1973).
Particle Structure
Particles are disrupted in 2% sodium phosphotungstate but in uranyl formate they are quasi-spherical and not penetrated by the stain (Fig. 8) (Jones & Mayo, 1973). The mean diameter of particles is 28 nm but particles have a broad distribution in the size range 25-32 nm suggesting heterogeneity in size. This was also suggested by the partial resolution of purified virus particle preparations into three bands by electrophoresis in polyacrylamide gels (Jones & Mayo, 1973).
Particle Composition
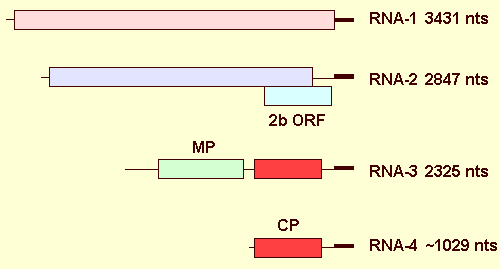
Nucleic acid: Virus particles contain several ssRNA species of which 3 (RNA-1, RNA-2, RNA-3) are genomic and up to 2 others (RNA-4, RNA-4a) are subgenomic. Particles contain c. 20% RNA (Jones & Mayo, 1973). Generally, the genomic RNA-3 and the subgenomic RNA-4 are present in much greater amount than the other species. The complete nucleotide sequences of the 3 genomic species of the Scottish isolate (Jones & Mayo 1973) have been determined; they contain 3,431 nt (RNA-1; accession no. U57047), 2,847 nt (RNA-2; accession no. U34050) and 2,325 nt (RNA-3; accession no. U57048). The subgenomic RNA-4 is estimated to be 1,029 nt in length and contains a copy of the coat protein gene that is located at the 3' end of RNA-3. The subgenomic RNA-4a contains a copy of the 2b ORF and is 851 nt in length (Xin et al., 1998).
Protein: The Mr of the virus coat protein (CP) subunit is 25,000 estimated by electrophoresis in polyacrylamide gels but the putative translation product of the CP gene of accession no. U57048 contains 213 aa and has an Mr of 23,635. The coat protein contains a conserved arginine motif that is responsible for the binding of the protein in the activation of the genome of EMoV, other ilarviruses, and Alfalfa mosaic virus (Ansel-McKinney et al., 1996).
Genome Properties
The genome comprises RNA-1, RNA-2 and RNA-3. RNA-1 contains a single ORF that encodes a putative polypeptide (1a protein) of 119.5 kDa (Fig. 9). The polypeptide contains two domains with sequence characteristics of methlytransferase and NTP-binding activities. RNA-2 contains 2 ORFs; the putative product of the larger ORF (2a protein) is 90.3 kDa and contains a domain with sequence characteristics of an RNA polymerase (Fig. 9). The smaller 2b ORF begins within the larger ORF and extends towards the 3' terminus of the molecule. The putative product (2b protein) is 20 kDa (Fig. 9) and is expressed via the subgenomic RNA-4a (Xin et al., 1998). RNA-3 is bicistronic and codes for a putative movement (3a) protein of 31.5 kDa and the coat (3b) protein of 23.6 kDa . The coat protein gene is expressed via the subgenomic RNA-4.
Relations with Cells and Tissues
No information.
Notes
The virus is distinct from Elm mosaic virus (a strain of Cherry leaf roll virus (Jones, 1985)) found in North America. The two viruses are unrelated serologically, belong to different virus families, and differ in particle properties, experimental host range and symptomatology (Jones, 1974, 1985). EMoV is serologically related to other members of subgroup 2 of the genus Ilarvirus (Roosinck et al., 2000). In particle properties and composition the virus resembles species in the other subgroups of the genus Ilarvirus but is distinguished from them serologically (Jones, 1974; Roosinck et al., 2000).
Acknowledgements
Photographs: Scottish Crop Research Institute, except Fig. 1 which is by courtesy of Dr K. Schmelzer.
Figures
References list for DPV: Elm mottle virus (404)
- Ansel-McKinney, Scott, Swanson, Xin & Gehrke, The EMBO Journal 15: 5077, 1996.
- Jones, CMI/AAB Descriptions of Plant Viruses 139: 1974.
- Jones, AAB Descriptions of Plant Viruses 306: 1985.
- Jones & Mayo, Annals of Applied Biology 75: 347, 1973.
- Roosinck et al., in Virus Taxonomy. Seventh Report of the International Committee on Taxonomy of Viruses, p. 923, eds M. V. H. Van Regenmortel et al., New York, San Diego. Academic Press, 2000.
- Schmelzer, Phytopathologische Zeitschrift 64: 39, 1969.
- Schmelzer, Schmidt & Schmidt, Archiv für Forstwesen 15: 107, 1966.
- Scott, Zimmerman & Ge, Archives of Virology 143: 1187, 1998.
- Scott, Zimmerman & Ge, Archives of Virology 148: 2063, 2003.
- Uyeda & Mink, Phytopathology 73: 47, 1983.
- Xin, Ji, Scott, Symons & Ding, Journal of Virology 72: 6956, 1998.